Research Article
Association of IL-17RC rs708567 with Systemic Lupus Erythematosus
1Molecular Medicine Center, Medical University-Sofia, 2 Zdrave Street, 1431 Sofia, Bulgaria
2Department of Clinical Laboratory and Clinical Immunology and Department of Nephrology, Medical University-Sofia, 1 Georgi Sofijski Street, 1431 Sofia, Bulgaria
3Hopitaux Drôme Nord, 607 avenue Geneviève de Gaulle-Anthonioz, 26102 Romans-sur-Isère, France
4Department of Medical Chemistry and Biochemistry, Medical University -Sofia, 2 Zdrave Street, 1431 Sofia, Bulgaria
5Department of Dermatology and Venereology, Medical University-Sofia, 1 Georgi Sofijski Street, 1431 Sofia, Bulgaria
*Corresponding author: Zornitsa Kamenarska, Molecular Medicine Center, Medical University-Sofia, 2 Zdrave Street, 1431 Sofia, Bulgaria, E-mail: kamenarska@yahoo.com
Received: July 3, 2018 Accepted: July 13, 2018 Published: July 18, 2018
Citation: Kamenarska Z, Hristova M, Dzhebir G, et, al. Association of IL-17RC rs708567 with Systemic Lupus Erythematosus. Madridge J Dermatol Res. 2018; 3(1): 65-67. doi: 10.18689/mjdr-1000114
Copyright: © 2018 The Author(s). This work is licensed under a Creative Commons Attribution 4.0 International License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
Background: Polymorphisms in the cytokine genes and their receptors are thought to influence the predisposition to systemic lupus erythematosus (SLE).
Objective: The aim of this case-control study was to investigate the association between the IL-17RC rs708567 polymorphisms and the susceptibility to SLE in Bulgarian patients. Altogether 44 patients with SLE as well as 93 unrelated healthy controls were included in this study.
Results: There is no difference of the allele and genotype distribution between the patients with SLE and the healthy controls. However, the CC genotype (p=0.03, OR 9.7, 95%CI 1.1-84) and the C allele (p=0.05, OR 2.4, 95% CI 1-5.8) appeared associated with arthritis. The CC genotype (p=0.002, OR 10.3, 95%CI 2.1-48.7) and the C allele (p=0.0008, OR 5, 95%CI 1.9-12.9) appeared associated with hematological diseases.
Conclusion: Our results indicate that the IL-17RC rs708567 polymorphism might play a disease modifying role in the development of SLE.
Keywords: IL-17RC; rs708567; Systemic Lupus Erythematosus.
Introduction
Systemic lupus erythematosus (SLE) is an autoimmune disease with multiple organ involvements and unknown etiology. However, the dysregulation of cytokine production or action is thought to have an important role in its development [1].
Interleukin 17 is a pro-inflammatory cytokine of the adaptive immune system, produced primarily by a new T helper cell subset termed ‘Th17'. Evidence indicates that the production of IL-17 is abnormally high in patients with SLE [2] and correlate with SLE disease activity [3, 4]. Furthermore, the frequency of Th17 is increased in the peripheral blood of patients with SLE [3, 5-7].
IL-17 exerts its function through the IL-17 receptor (IL-17R) complex. An integral part of it is IL-17 receptor C (IL-17RC) - a single-pass type I transmembrane protein, encoded by the IL-17RC gene [7]. Being an essential part of the IL-17/IL-17R signaling axis, the IL-17R complex mediates the signal transduction of the IL-17 cytokine family which promotes the production of the pro-inflammatory cytokines such as tumour necrosis factor α (TNF-α), interleukin-1β (IL-1β), and interleukin-6 (IL-6). Current evidence indicates that both the dysfunction of IL-17RC and the IL-17/IL-17R signaling axis are implicated in a number of human diseases. The SNP rs708567 C/T is a common missense mutation (S111L) within the IL-17RC gene which may play an important role in impairing IL-17RC gene function [4].
The objective of our study was to determine whether the IL-17RC rs708567 polymorphism is a risk factor for the development of SLE in Bulgarian patients and to define its contribution to the increased risk.
Patients and Methods
Clinical Material
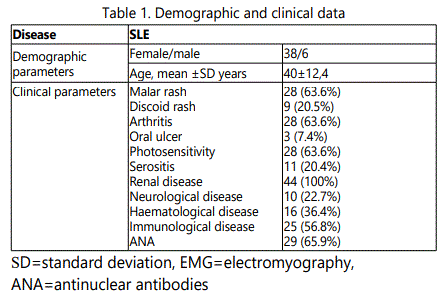
Forty-four patients with SLE were included in this study, 38 of them were female and 6 - male. The mean age was 40 with a range of 15 to 78 years. The patients have been followed for a mean of 10 years at the Department of Dermatology and Venereology and at the Department of Nephrology, Medical University of Sofia. All the patients with SLE met the American College of Rheumatology (ACR) criteria [8].
Ninety-three anonymous non-related healthy individuals, matched for sex, age and ethnicity with the patients were included for genetic analysis. They were selected from the Bio Bank of the Molecular Medicine Center and National Genetic Laboratory.
Genetic Analysis
The scientific investigation presented in this article has been carried out in accordance with The Code of Ethics of the World Medical Association (Declaration of Helsinki) for experiments involving humans. The study was approved by the local ethics committee at the Medical University- Sofia. All participants signed informed consent and venous blood was drawn for DNA isolation. Genomic DNA was extracted from the peripheral blood with the Chemagen DNA purification kit, using Chemagic Magnetic Separation Module I (Chemagen AG). The analysis of IL-17RC rs708567 was performed by using TaqMan genotyping assay.
Statistical Analysis
Allele and genotype frequencies were compared between DM and SLE cases and controls, using Fisher's exact test to calculate p values for 2 × 2 tables. Where significant, data were expressed as p-value, odds ratios (OR) with exact 95% confidence intervals (CI). Test for Hardy-Weinberg equilibrium was done by χ2 statistics (SPSS 22, Inc. Chicago, Illinois, USA).
Results
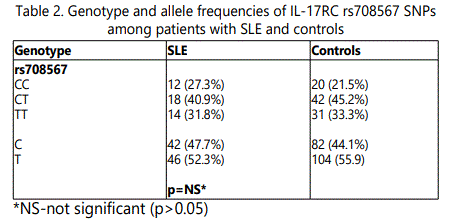
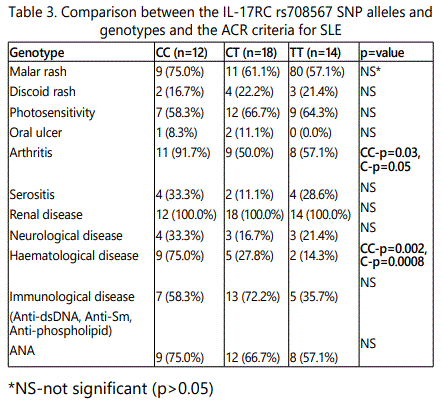
The distribution of the genotypes was in Hardy-Weinberg equilibrium. There is no difference of the allele and genotype distribution between the patients with SLE and the healthy controls. However, the CC genotype (p=0.03, OR 9.7, 95%CI 1.1-84) and the C allele (p=0.05, OR 2.4, 95% CI 1-5.8) appeared associated with arthritis. The CC genotype (p=0.002, OR 10.3, 95%CI 2.1-48.7) and the C allele (p=0.0008, OR 5, 95%CI 1.9-12.9) appeared associated with hematological diseases.
Discussion
Animal models have demonstrated that blockade of IL-17 decreases lupus manifestations [9]. On the other hand, SLEderived B cells have been shown to increase anti-DNA production when cultured in the presence of IL-17 [10].
The rs708567 polymorphism is located in the exon 4 of the IL-17RC gene (G/A +6313), but its functional effect is indefinite. Earlier studies could not demonstrate any relationship between the IL-17RC rs708567 SNP and IL-17 plasma levels [11].
Studies found the SNP associated with the development of scoliosis [12, 13] and malaria [14] but it had no role for the development of rheumatoid arthritis [11]. We found no association of that polymorphism with the development of SLE, which means that the SNP probably has no role in the pathogenesis of the autoimmune diseases. However, we found it associated with haematological disease and arthritis. Earlier, it was demonstrated that the IgM-rheumatoid factor and anti-CCP antibodies were significantly less frequent in patients with RA carrying the IL-17RC*T/T genotype than those carrying *C/C and *C/T genotypes. All these results demonstrate the disease modifying role of the IL-17RC rs708567 SNP.
Conclusion
This is the first study to investigate the role of the 17RC rs708567 SNP in the pathogenesis of SLE. The analysis should be repeated in a larger cohort in order to confirm our preliminary results.
Acknowledgements
The study was fulfilled with the financial support of Medical University-Sofia under contract D-67/2016.
Conflict of Interest
The authors declare that they have no conflict of interest.
References