Research Article
Categorization of Metalloboranes using Skeletal Numbers
Kironde Road, Kyadondo Block 224, Plot 5387, Kampala, Uganda
*Corresponding author: Enos Masheija Rwantale Kiremire, Professor, Kironde Road, Kyadondo Block 224, Plot 5387, Kampala, Uganda, E-mail: kiremire15@yahoo.com
Received: February 20, 2019 Accepted: February 25, 2019 Published: March 1, 2019
Citation: Kiremire EM. Categorization of Metalloboranes using Skeletal Numbers. Int J Chem Res. 2018; 1(1): 24-34. doi: 10.18689/ijcr-1000105
Copyright: © 2019 The Author(s). This work is licensed under a Creative Commons Attribution 4.0 International License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
Metalloboranes have been analyzed and categorized using skeletal numbers. Skeletal numbers are extremely helpful in categorizing chemical clusters literally ranging from a single to an infinite number of skeletal elements of the main group and transition metals. The method involves decomposing a cluster formula comprising of a number of skeletal elements using skeletal numbers into a single whole number (K). Knowing the number of skeletal elements in a cluster (n), the K(n) parameter is obtained. The type of series S=4n+q is readily derived from the K(n) parameter. This is followed by the derivation of the capping parameter, Kp=CyC[Mx] and the categorization parameter K*=Cy+Dz where y+z=n, the number of skeletal elements in the cluster. The cluster valence electrons can be calculated from the relationship VE=12y+14z+2 for transition metal clusters or VE=2y+4z+2 for main group elements or VE=2y+4z+2+10m where m=the number of transition metal elements present when a cluster has a mixture of elements from the main group and transition metals. The categorization parameter can also act as a guide to predict a possible symmetry of a cluster and to construct an isomeric skeletal shape of a cluster. This approach was used to categorize more than 20 metalloborane clusters as well as a few non-metalloborane clusters for comparison purposes. The isomeric skeletal structures so constructed obey the 8 or 18 electron rule as defined by coordination theory.
Keywords: Capping theory; Capping parameter; Categorization parameter; Coordination theory; Skeletal number; 4N series.
Introduction
In chemistry, some clusters have been categorized as closo, nido, arachno, hypho and klado, among others [1-7]. In this regard, the polyhedral skeletal electron pair theory (PSEPT) has been extremely useful in this process [8-11]. On closer analysis, it was observed that the transition metal clusters follow the series S=14n+q while the main group element clusters follow a parallel series S=4n+q where n represents the number of skeletal elements in a cluster and q is a numerical determinant that determines the category of the cluster. For categorization of clusters, it was found to be more convenient to utilize the S=4n+q for both main group and transition metal clusters except when the cluster valence electrons are being calculated whereby S=14n+q is applied for transition metal clusters [3,4]. The skeletal numbers which were derived from the series have been found to be extremely useful in the derivation of chemical graph theory and equations for calculating cluster valence equations [6]. In addition, they can be used as a guide to predict the possible geometrical and graphical structures of clusters [6]. Furthermore, skeletal numbers can be regarded as individual contributions to the skeletal cluster number parameter K(n) by the elements working together as a ‘team’ in the cluster. The 4N series approach was earlier applied to categorize metalloboranes. In this paper, the application of skeletal numbers to categorize metalloboranes will be demonstrated.
Results and Discussions
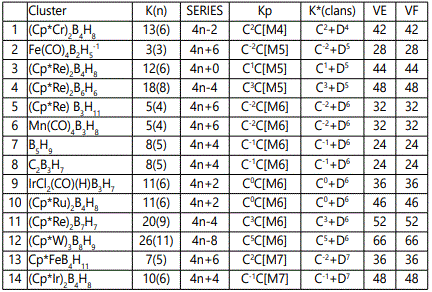
The application of the skeletal number to categorizing clusters is a new concept [5] and the K(n) parameter is usually a whole number. Since this approach is new, the skeletal numbers have been reproduced in table 1.
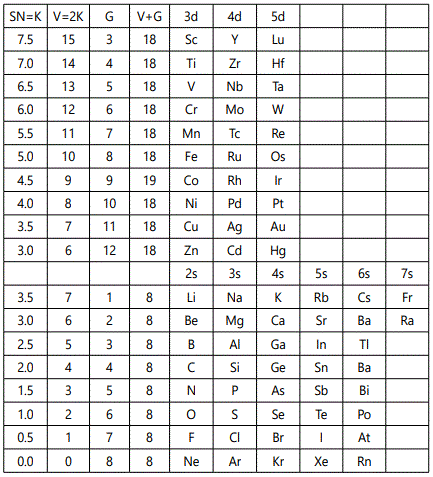
SN: Skeletal Number; V: Valence; G: Group valence electrons.
The Condensed outline of the Method for categorization of Clusters
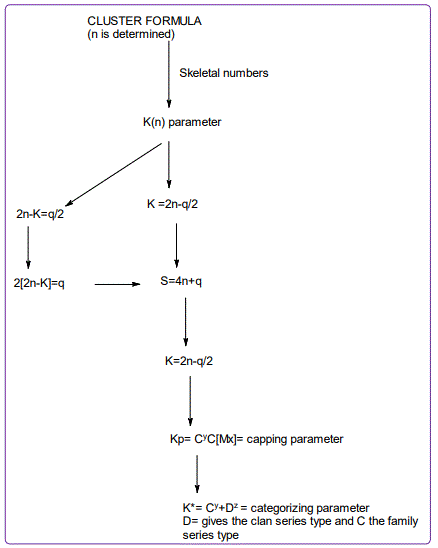
The first step in cluster categorization is to convert the cluster formula into a number. This approach focuses on the compounds or clusters from the main group or transition metals. This is now possible since each of the periodic group of elements was assigned a skeletal number as well as the ligands. The determination of the K value for a ligand is simple; LIGANDS: FOR EVERY 1 ELECTRON DONATED, K=-0.5 AND FOR EVERY 1 ELECTRON REMOVED, K=+0.5. So the cluster number K is readily calculated for any cluster with or without ligands. This is summarized in Scheme 1 and illustrated by in the following examples and worked out exercises most of them with detailed explanations.
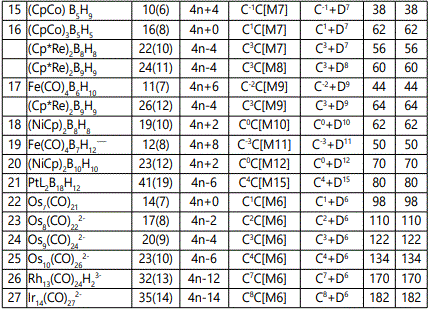
In the examples 1-7, the use of skeletal numbers to categorize clusters has been highlighted. A cluster is categorized into a CLAN series Dz and the family series Cy via the categorization parameter K*= =Cy+Dz. The same parameter can be used to calculate the cluster valence electrons VE=12y+14z+2 for transition metals and VE=2y+4z+2 for main group elements. The same parameter can be used as a guide to predict the shape of the cluster. For instance, K*=C0+D6, is an octahedral symmetry, K*=C1+D6 is a mono-capped octahedron, K*=C2+D6; bi-capped octahedron, K*=C3+D6; is a tri-capped octahedron and so on. Also a list of worked out examples mainly metabollanes from 1 to 22 were analyzed and categorized into clans and families and are given in table 2. Their graphical skeletal structures constructed according to skeletal valence principle: while they may involve inflated cluster valence electrons, they generate the correct cluster formulas, they act as a guide for predicting possible structure of the cluster, and above all, the capping cluster formula which is so accurate in determining cluster valence electrons, is based on the same principles.
Coordination Theory
According to the 4N series approach to clusters it makes understanding much easier if we regard each linkage line to a skeletal element(node) as donating 1 electron to the nodal point. In that way, methane, CH4 can be explained as the central carbon atom which has its own 4 electrons receiving donations from the 4H atoms in order to attain the 8-electron configuration. Each of the 4 hydrogen ligands has donated 1 electron to the node and the octet rule has been obeyed. This is the way coordination theory is viewed according to the 4N series approach. This is illustrated in EX-1A. However if we consider BH5, according to the 4N series approach, the node(B) has 3 valence electrons and the addition of 5 electrons to the node will give the node a total of 8 electrons enabling the node to achieve the eighteen electron rule. In fact, the cluster BH5 appears in a disguised form as BH4—.According to the4N series method, there is no difference between a charge of -ve one and 1H ligand donor.
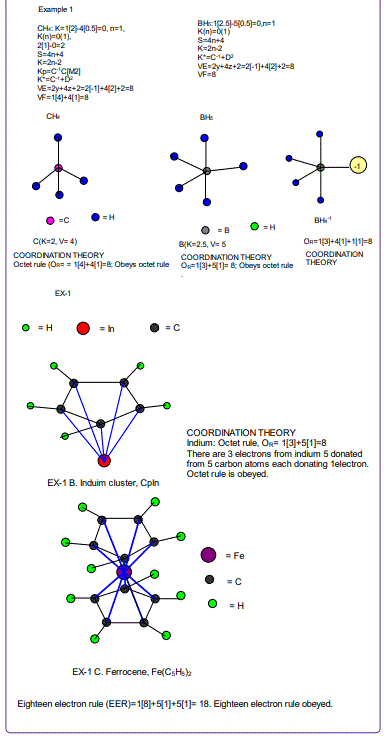
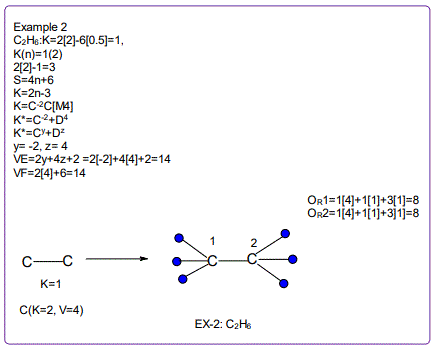
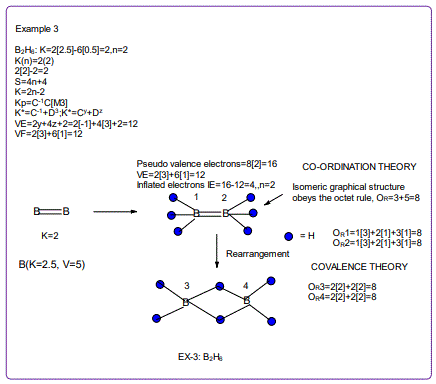
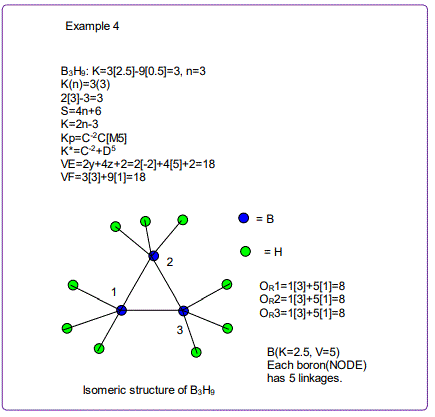
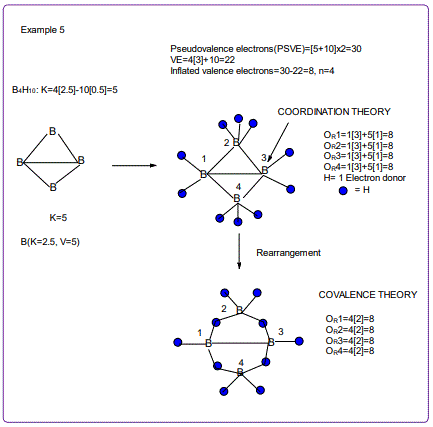
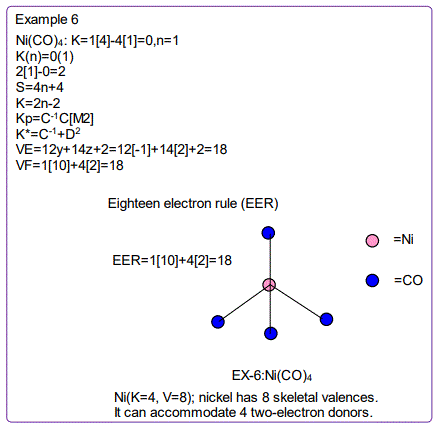
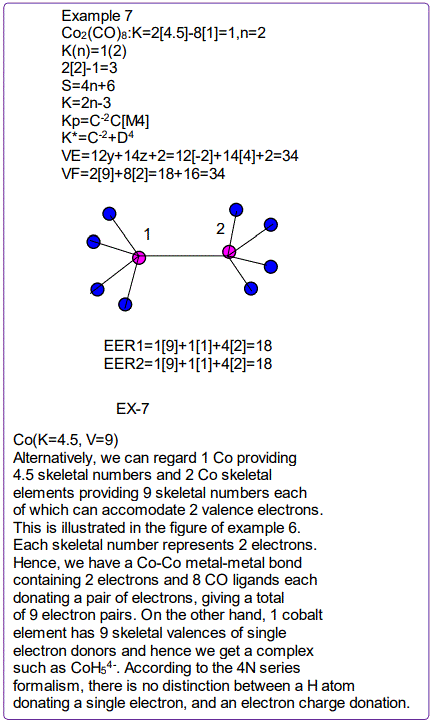
All the isomeric graphical structures are constructed in line with coordination theory and they obey the octet rule for main group elements or eighteen electron rules for transition metals.
Categorization of Clusters into Clans and Families
The sample of clusters mainly metaboranes were categorized into clans (Dz) and families (Cy). As can be seen from table 2, they range from D4 to D15 and with families from C-2 to C5. The following clan series of the metalloboranes were identified, D4(1), D5(3), D6(8), D7(4), D9(1), D10(1), D11(1), D12(1) and D15(1). Also some D6 samples were added for comparative purposes. They belong to the family series C1, C2, C3, C4, C7 and C8 of the D6 clan. The isomeric skeletal shape is also provided with explanation for each of the clusters analyzed and categorized. The corresponding isomeric skeletal structures are from figures 1-24. The categorization symbol K*=Cy+Dzcan be viewed as representing a nucleus, Dz and the outer shell Cy. The valence electrons of the cluster comes from the nuclear component which follow the VE1=S=14z+2 while the capping electrons are given by VE2=12y giving the total valence electrons VE=VE2+VE1=12y+14z+2.
The added D6 clan series have families ranging from C1 to C8. The isomeric graphical skeletal structures of clusters adopted in this paper are constructed based on the simple coordination theory as defined in this work. Once such a structure has been constructed, the original cluster formula usually emerges from it. Furthermore, each node obeys the 8-electron or 18-electron rule if it is from the main group or transition metals respectively. Each of the linkage in the cluster represents a 1-electron donor to the node (vertex). Each node is linked to the number of lines in accordance with the skeletal valence of the fragment at the vertex. The decomposition of a cluster formula into a number followed by the categorization of the cluster into clan and family type (K*=Cy+Dz) is well explained in each of the 22 cluster examples selected.
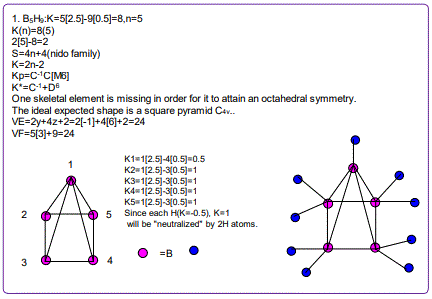
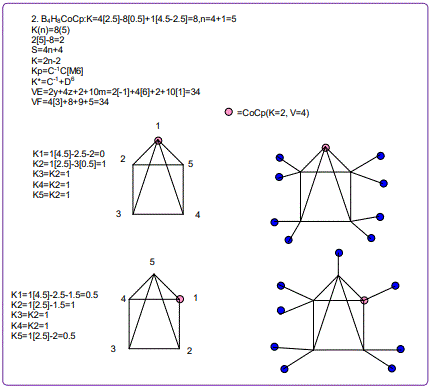
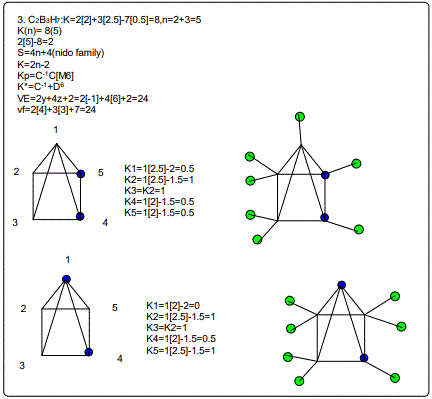
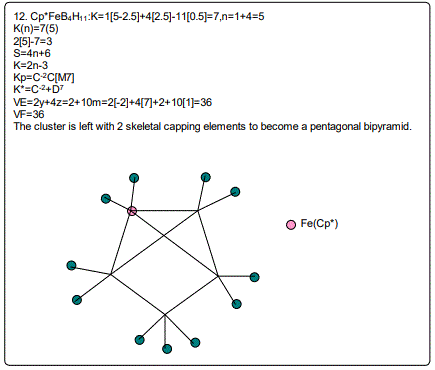
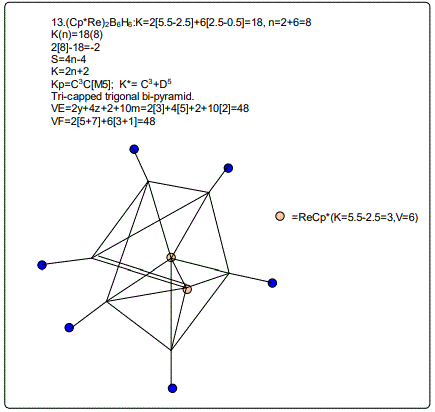
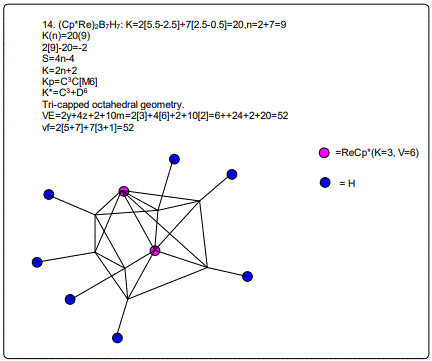

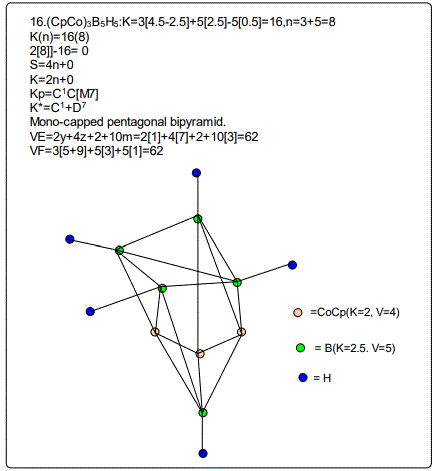
18. (CpCo)B5H9:K=1[4.5-2.5]+5[2.5]-9[0.5]=10,n=1+5=6
K(n)=10(6)
2[6]-10=2
S=4n+4(nido)
K=2n-2
Kp=C-1C[M7]
K*=C-1+D7
VE=2y+4n+2+10m=2[-1]+4[7]+2+10[1]=38
VF=1[5+9]+5[3]+9[1]=38
The isomeric structure has not been provided as it can be generated in the same procedures as above ones.
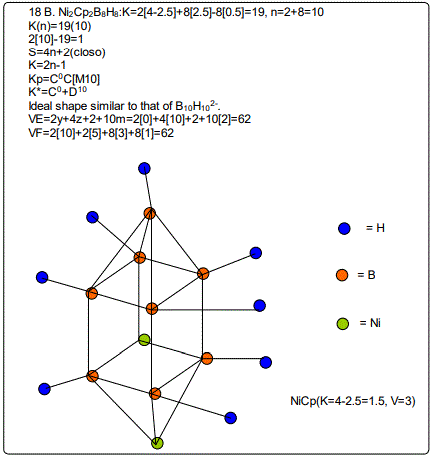
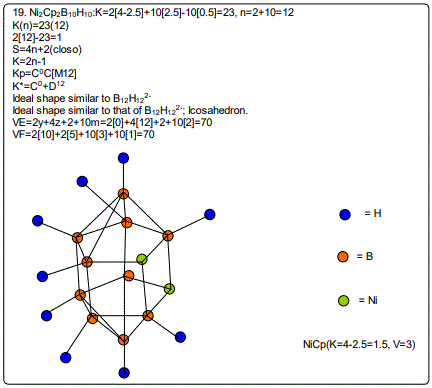
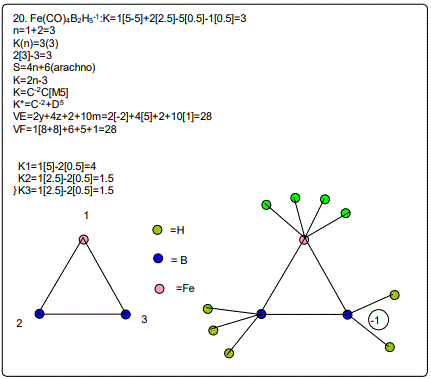
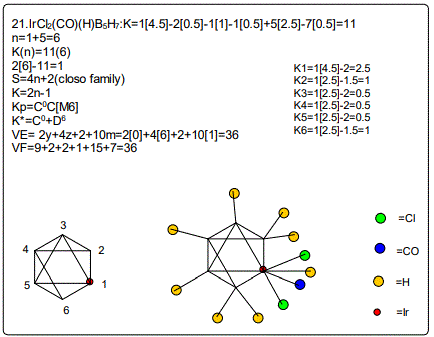
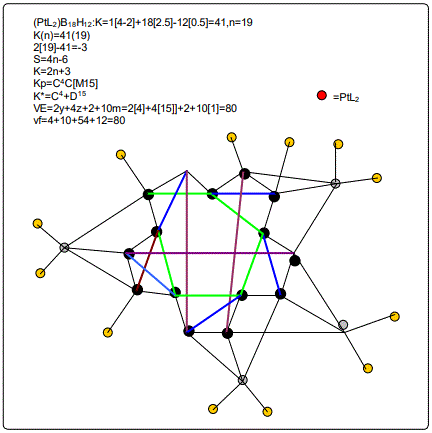
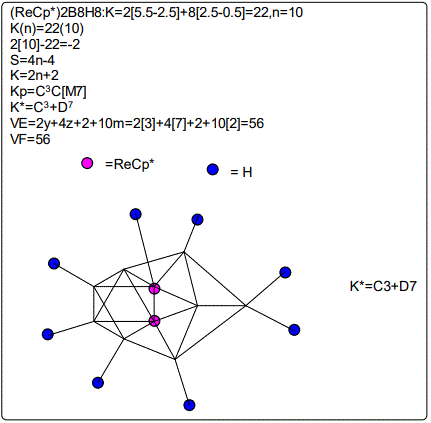
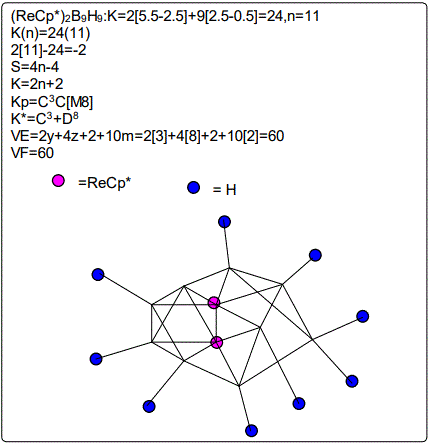
Derivation of Information from the Categorization Parameter K*=CY+DZ
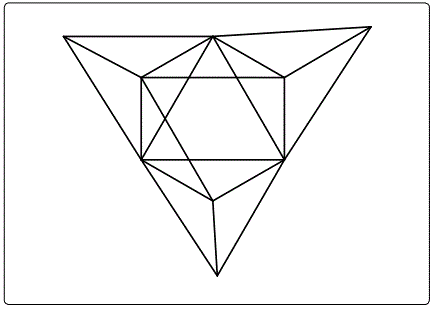
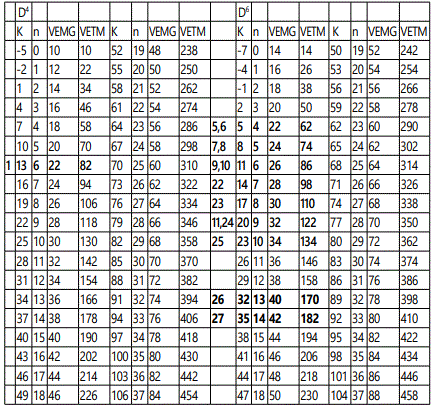
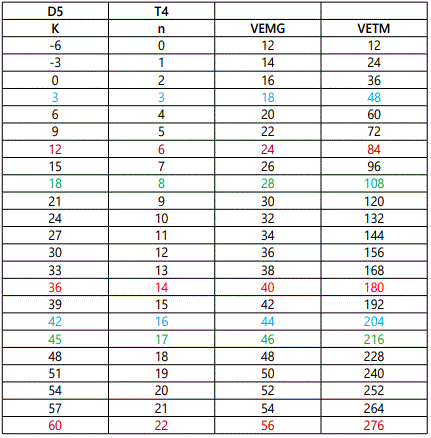
Once a cluster formula has been converted into a categorization parameter K*=Cy+Dz all the important information is concealed into that symbol. All that is needed to know is whether it refers to the main group or transition metals. For instance, K*=C3+D6; refers to a cluster with 9 skeletal elements (n=3+6=9), VE=12y+14z+2=12[3]+14[6]+2=122 (transition metals) and VE=2y+4z+2=2[3]+4[6]+2=32 (main group elements). The ideal shape will be a tri-capped octahedron as shown in figure 25. The clan series do generate an infinite range of K(n) and VE series. The numerical series from n=0 to n=37 for D4 and D6 for both main group and transition metals are given in table 3. In order to highlight the significance of categorization of clusters into clans, a small sample of D5 series was categorized and is given in the examples G1 to G6. The final categorization results of the D5 series are: K*=C5+D5; C9+D5; C10+D5; C11+D5; C12+D5; and C17+D5. Also selected clan series of D5 are given in table 4. The positions of the K(n) and VE values of G1 to G6 clusters are highlighted in table 4. The D5 and D6 represent clan clusters whose structures are centered around an ideal trigonal bipyramid and octahedral nuclei respectively. The ideal structures are shown in figure 26.

The variation of the skeletal number in similar cluster formulas
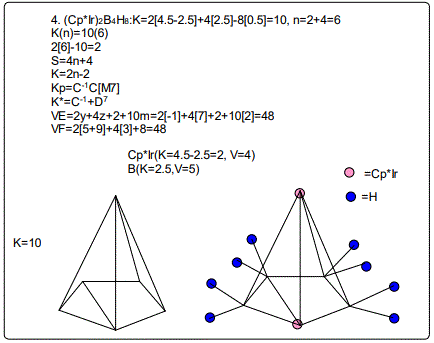
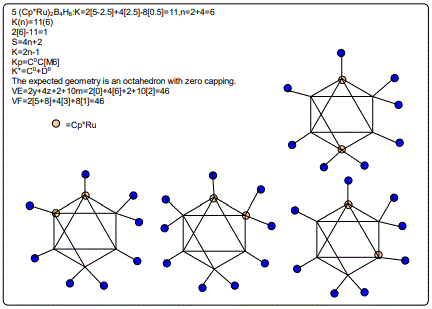
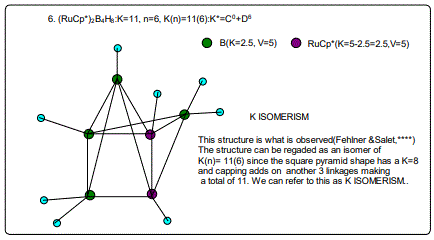
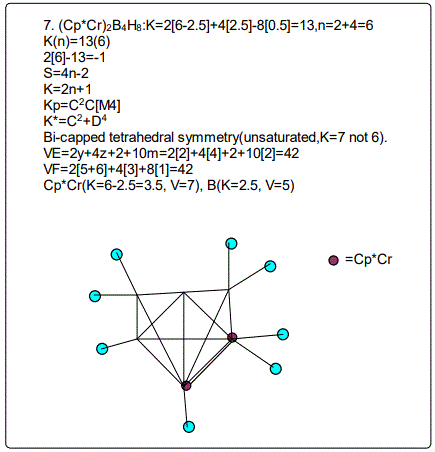
Let us focus on the case of the following clusters (Cp*M)2B4H8, M=Ir, Ru, Re and Cr as an illustration of the concept. The corresponding K(n) parameter and categorization parameter K* are as follows;10(6),C-1+D7; 11(6),C0+D6; 12(6),C1+D5;13(6),C2+D4. The K values for Ir, Ru, Re, and Cr are 4.5, 5.0, 5.5 and 6 respectively the K(n) parameter starting with K=10 to 11, 12 and then 13 for the same number of skeletal elements n=6. As can be seen, the number of skeletal elements remains the same while the value of the cluster number K increases. This results into the increase in the capping index while the nuclear index is decreasing. In the parameter K*=Cy+Dz and n=y+z. Since n is constant, the increase in y will naturally result in the corresponding decrease in z. Hence, the increase in the K value results in the increase of the capping index (the outer shell expands) and a decrease in the nuclear index (nucleus shrinks).
G1. F=Os10(C)(CO)24:K=10[5]-1[2]-24[1]=10,n=10
K(n)=24(10)
S=4n-8
K=2n+4
Kp=C5C[M5]
K*=C5+D5
K*=Cy+Dz
VE=12y+14z+2=12[5]+14[5]+2=132
VF=10[8]+1[4]+24[2]=132
G2. Rh14(CO)254-:K=14[4.5]-25[1]-4[0.5=36,n=14
K(n)=36(14)
S=4n-16
K=2n+8
Kp=C9C[M5]
K*=C9+D5
K*=Cy+Dz
VE=12y+14z+2=12[9]+14[5]+2=180
VF=14[9]+25[2]+4[1]=180
G3. Rh15(CO)273-:K=15[4.5]-27[1]-3[0.5]=39,n=15
K(n)=39(15)
2[15]-39=-9
S=4n-18
K=2n+9
Kp=C10C[M5]
K*=C10+D5
K*=Cy+Dz
VE=12y+14z+2=12[10]+14[5]+2=192
VF=15[9]+27[2]+3[1]=192
G4. Pd16(CO)13L9:K=16[4]-13[1]-9[1]=42,n=16
K(n)=42(16)
2[16]-42=-10
S=4n-20
K=2n+10
Kp=C11C[M5]
K*=C11+D5
VE=12y+14z+2=12[11]+14[5]+2=204
VF=16[10]+13[2]+9[2]=204
G5. Rh17(CO)303-: K=17[4.5]-30[-1]-3[0.5]=45,n=17
K(n)=45(17)
2[17]-45=-11
S=4n-22
K=2n+11
Kp=C12C[M5]
K*=C12+D5
VE=12y+14z+2=12[12]+14[5]+2=216
VF=17[9]+30[2]+3[1]=216
G6. Rh22(CO)374-:K=22[4.5]-37[1]-4[0.5]=60,n=22
K(n)=60(22)
2[22]-60=-16
S=4n-32
K=2n+16
Kp=C17C[M5]
K*=C17+D5
VE=12y+14z+2=12[17]+14[5]+2=276
VF=22[9]+37[2]+4[1]=276
Conclusion
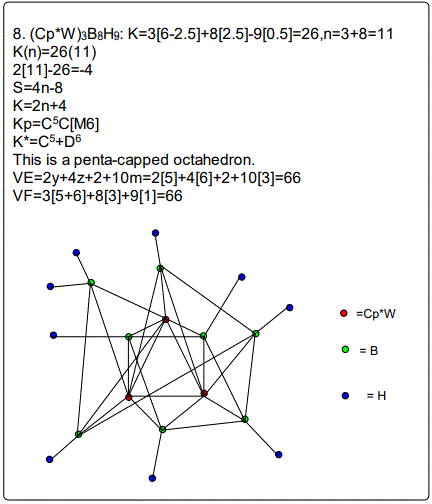
A good range of metalloboranes have been analyzed and categorized into clans and families of clusters using skeletal numbers. The cluster valence electrons and isomeric skeletal structures can readily be derived. The metalloborane (Cp*W)3B8H9 with a categorizing parameter K*=C5+D6 was found surprisingly highly capped unlike other metalloboranes. A formula for calculating cluster valence electrons for a cluster with a mixture of non-metals was derived namely: VE=2y+4z+2+10m where y is a capping index for Cy elements and z is a capping index for Dz elements and m is the number of metal skeletal elements in the cluster. Thus in the case of the (Cp*W)3B8H9, y=5,z=6 and m=3 which gives us a cluster valence value of 66. The 10m component of the formula is for metal adjustment to take into account the additional cluster valence electrons introduced by metal atoms present.
Acknowledgement
I wish to thank Dr. Penelope Emma Kiremire for editing the draft paper.
References
- Greenwood NN. Metalloboranes. Pure Appl Chem. 1983; 55(9): 1415-1430. doi: 10.1351/pac198355091415
- Housecroft CE, Sharpe AG. Inorganic Chemistry. 2nd Edition. Pearson, Prentice Hall, Harlow, England. 2005.
- Kiremire EM. The Application of the 4n Series Method to Categorize Metalloboranes. Int J Chem. 2016; 8(3): 62-73. doi: 10.5539/ijc.v8n3p62
- Kiremire EM. Classification of Zintl Ion Clusters Using 4n Series Approach. Orient J Chem. 2016; 32(4): 1731-1738. doi: 10.13005/ojc/320401
- Kiremire EM. The Outstanding Applications of Skeletal Numbers to Chemical Clusters. Int J Chem. 2017; 9(3): 28-48. doi: 10.5539/ijc.v9n3p28
- Kiremire EM. The Capping Theory of Chemical Clusters Based on 12N/14N Series. Int J Chem. 2018; 10(4): 130-154. doi: 10.5539/ijc.v10n4p130
- Miessler GL, Fischer PJ, Tarr DA. Inorganic Chemistry. 5th Edition, Pearson Education, Inc., Upper Saddle River. 2013.
- Mingos DMP. A General Theory for Cluster and Ring Compounds of the Main Group and Transition Elements. Nature Physical Science. 1972; 236: 99-102. doi: 10.1038/physci236099a0
- Wade K. The structural significance of the number of skeletal bonding electron-pairs in carboranes, the higher boranes, and borane anions and various transition metal carbonyl cluster compounds. Chem Commun. 1971; 792-793. doi: 10.1039/C29710000792
- Wade A. Structural and Bonding Patterns in Cluster Chemistry. Adv Inorg Chem Radiochem. 1976; 18: 1-66. doi: 10.1016/S0065-2792(08)60027-8
- Welch AJ. The significance of Wade’s rules. Chem Commun. 2013; 49: 3615-3616. doi: 10.1039/C3CC00069A


