Research Article
Proteomic analyses of Urine Exosomes reveal New Biomarkers of Diabetes in Pregnancy
1Department of Medicine, Biomarkers Laboratory, O'Brien Center for Acute Kidney Injury Research, UC San Diego, USA
2Department of Pediatrics, Center for Promotion of Maternal Health and Infant Development, UC San Diego, USA
3Department of Reproductive Medicine, UC San Diego, USA
4Clinical and Translational Research Institute, UC San Diego, USA
5Department of Chemistry & Biochemistry, Biomolecular & Proteomics Spectrometry Facility, UC San Diego, USA
6Departments of Medicine, Pathology and Pediatrics, UC San Diego, USA
*Corresponding author: Satish P Ramachandra Rao, Department of Medicine, Biomarkers Laboratory, O'Brien Center for Acute Kidney Injury Research, UC San Diego, USA, Tel: +91-8152-243138, Fax:+91-8152-243008, E-mail: satishrao@sduu.ac.in; satishrao@ucsd.edu
Received: January 8, 2016 Accepted: February 1, 2016 Published: February 1, 2016
Citation: Ramachandrarao SP, Hamlin AA, Awdishu L, et al. Proteomic analyses of Urine Exosomes reveal New Biomarkers of Diabetes in Pregnancy. Madridge J Diabetes. 2016; 1(1): 11-22. doi: 10.18689/mjd-1000103
Copyright: © 2016 The Author(s). This work is licensed under a Creative Commons Attribution 4.0 International License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
Objective: To evaluate 24 hour urine exosome protein content changes among pregnant US subjects with diabetes and obesity during early pregnancy.
Methods: The exosome proteome content from 24 hour urine samples of pregnant subjects with gestational diabetes mellitus (GDM, N=8) and pre-gestational Type 2 diabetes (PGD, N=10) were compared with control samples (CTRL, N=10) obtained at week 20 of pregnancy. Differences in exosome protein load between groups was identified by liquid chromatography/mass spectrometry, analyzed by linear regression in negative binomial distribution, visualized in MetaboAnalyst (version 3.0), and validated by western immunoblotting.
Results: At the 20th week of pregnancy, we identified 646, 734 and 856 proteins in exosomes from 24 hour urine samples of patients from the CTRL, GDM and PGD groups, respectively. S100 calcium binding protein A9, damage associated molecular pattern (DAMP) signal, was found to be significantly increased in both GDM and PGD subjects. In GDM subjects the peptide counts for S100A9 protein independently correlated with maternal obesity and macrosomia of the newborn infants. Early to late pregnancy developmental changes in the GDM group were shown to utilize pathways and protein expression levels differently from those in PGD or CTRL groups.
Conclusions: Urinary exosome proteomic analysis non-invasively provides insights into maternal changes during diabetic pregnancy. Exosome biomarkers early in pregnancy can be potentially used to better understand pathophysiologic mechanisms of diabetes at a cellular level, and to distinguish between gestational and pre-gestational diabetes at the pathway level. This information can aid intervention efforts to improve pregnancy outcomes in women with diabetes.
Keywords: Damage associated molecular pattern; Diabetic pregnancy; Exosome; Gestational diabetes; Urine exosomes; Proteomics; S100A9.
Abbreviations: PGD: Pre-gestational diabetic; MVBs: Multivesicular bodies; PLS-DA: Partial-Least Squares Discriminant Analysis; VIP: Variable Importance in Projection; FDR: False Discovery Rate; PLSD: Partial Least Squares Discriminant; DAMP : Danger-Associated Molecular Pattern.
Introduction
Diabetes during pregnancy increases the risk of poor pregnancy outcomes. Poorly controlled maternal diabetes may increase the risk of malformations involving multiple organ systems, such as fetal cardiac and spinal abnormalities during the first trimester and even fetal loss during the third trimester [1]. Renal glomerular filtration rate and plasma flow increase during normal pregnancy, leading to increases in proteinuria, glucosuria, and lower serum osmolality and sodium levels [2]. In pregnancies complicated by diabetes these changes may be exacerbated causing subclinical renal insufficiency in addition to the other unfavorable outcomes mentioned [3]. Pre-gestational diabetic (PGD) mice develop accelerated renal pathology [4], suggesting that stimuli that elicit benign responses in the non-diabetic pregnancies may lead to renal dysfunction or injury in diabetic pregnancies [5,6]. Further, as demonstrated by the collaborative perinatal project, the PGD mothers carry different risks of giving birth to malformed infants than do the GDM mothers [7]. Mechanisms underlying end organ damage or malformation and the maternal/fetal metabolic programing affecting the kidney function of the newborn still remain incompletely defined [8-11]. New tools that are noninvasive yet specific enough to examine the pathologic alterations in maternal diabetes during pregnancy are required to study clinically relevant sequelae specific to GDM or PGD phenotypes. We believe that urinary exosomes represent a noninvasive paradigm that potentially may be able to address phenotypespecific questions.
Exosomes are membrane bound biologically active nanovesicles released from every cell type [12,13]. They are formed by inward budding of late endosomes, producing Multivesicular bodies (MVBs). They are then released into the surrounding biofluid by fusion of MVBs with the plasma membrane [14]. The endocytic origin of exosomes and the cellular nature of their contents enable them to be recognized as a rich source of information on the status of the cells producing them. Because their contents significantly change in response to stress or injury [15] they are also potent mediators of intercellular communication [16]. Average human kidneys filter 1500-2000 liters of plasma on a daily basis to produce roughly 1.5 liters of urine. Together with the observation that urinary exosomes contain 3-5% of the total urinary proteins, exosomes represent enriched background information on the subject's systemic status.
Diabetic pregnancy is a stressor on multiple organs, including the kidney. Further, GDM is qualitatively a different type of stress compared to PGD [7]. Therefore, at an early time point during diabetic and normal pregnancy we sought to characterize urine exosome proteins from CTRL (n=10), PGD (n=10) and GDM (n=8) women. We also investigated pathway level differences between pre-gestational and gestational diabetes, as well as developmental changes from early to late pregnancy as reflected in their urine exosome proteomes.
Materias & Methods
Patient Selection
Singleton pregnancies who received prenatal care at the
University of California San Diego Medical Center from 2011
to 2013 were consented to participate in the study. The study
design and recruitment was reviewed and approved by the
institutional review board. Informed consent for participation
was obtained from all study participants. Inclusion criteria
included pre-existing Type 2 diabetes or gestational diabetes.
Diagnosis for gestational diabetes followed the criteria
established by the California Sweet Success Program. This
criterion includes at least one of the following: a first trimester
HbA1c between 5.7-6.4% (39-46 mmol/mol), a fasting plasma
glucose between 92-126 mg/dL, or a second trimester two
hour glucose tolerance test with a fasting plasma glucose of
greater than or equal to 92 mg/dL, one hour greater than or
equal to 180 mg/dL, ora two hour plasma glucose greater
than or equal to153 mg/dL [17]. Control samples were
obtained from consenting women who did not have preexisting diabetes and screened negative for gestational
diabetes. Study participants were required to collect a 24 hour
urine sample at 20 and 36 weeks gestation.
Demographics
Data was collected from a chart review. Information regarding diabetes management and control were obtained during first, second, and third trimesters, including HbA1c, fasting plasma glucose levels, diabetes medications (oral hypoglycemics and insulin), and any maternal medical complications. Fetal outcomes including ultrasound, estimated third trimester fetal weight, and birth weight were also collected.
Study design: Urine samples at the 20th week (early time point) of pregnancy from 8 GDM, 10 PGD and 10 CTRL subjects were used for exosome isolation and proteomic analysis in replicates. A subset of these subjects that completed sample collection during 36th week of pregnancy (5 GDM, 3 PGD and 4 CTRL pregnancy subjects) were separately analyzed to evaluate differences between the 20th and 36th week. Proteomic data were analyzed using negative binomial distribution as well as MetaboAnalyst (version 3.0) and validated by Western immunoblotting where applicable.
Urine Sampling and Processing
24 hour urine samples were collected spanning 2 days. The first void of Day 1 was not collected. All voids were collected from the second void on Day 1 to the first void on Day 2, including all nocturnal voids. During collection the samples were kept on ice or refrigerated. After 24 hours collected urine samples were centrifuged at 3000 x g for 30 min. The pH of the resulting supernatant was adjusted to 7.0, aliquoted, and frozen at -70 °C until further analysis.
Exosome preparation and proteomic analysis
Exosomes from frozen pregnancy urine samples were prepared using an in-house protocol developed based on the solvent exclusion principle using polyethylene glycol (PEG)- induced precipitation, as described in our recent publication [18]. One-dimensional SDS-PAGE of the exosome proteins prior to in-gel trypsinization was performed.
Data Processing and Statistical Analysis
In all analyses, p ≤0.05 was considered statistically significant. Log transformation of raw data were performed before the tests. Analyses including Student's t-tests, Partial-Least Squares Discriminant Analysis (PLS-DA) and Variable Importance in Projection (VIP) were performed with the MetaboAnalyst 3.0 (www.metaboanalyst.ca) web portal [19]. Negative binomial generalized linear regression models were fit using the statistical software R version 3.1.2.20
PLS-DA and VIP were used both for the classification and significant feature selection [19] with a False Discovery Rate (FDR) of ≤10 % to filter protein candidates for western immunoblotting validation. Negative binomial generalized linear models were fit to determine significant differences in protein counts between CTRL, GDM, and PGD subjects utilizing the raw count data and estimating gene-wide size factors and estimated dispersion-mean relationship via the methods outlined in Anders and Huber in the R package DESeq2 [21]. Significance was reported based on a filtered FDR-adjusted p-value.
Western Immunoblotting and Quantification
Antibody against S100 A9 was purchased from Proteintech Group, Inc., (Chicago, IL, USA). HRP-conjugated secondary antibody was from GE Life Sciences (Piscataway, NJ, USA). SDS-PAGE gels (with 10% acrylamide) were used to resolve 100 μg of protein from exosomes of normal and diabetic pregnancy urine samples. Immunoblotting and quantification with Image J software (NIH) was done using methods described in our previous publications [22] and plotted using Graph Pad Prism software (San Diego, CA, USA).
Results
Patient Demographics Show Significant Differences between
Diabetic and Non-Diabetic Pregnancy Subject Phenotypes
We studied 24 hour urine samples from 28 mothers: 10
non-diabetic subjects that served as controls (CTRL) and 18
with diabetes (8 gestational, GDM and 10 pre-gestational,
PGD) recruited from UC San Diego Reproductive Medicine
clinics. Clinical disease data on 6 GDM, 7 PGD and 6 CTRL
mothers was collected. Newborn birth weight in PGD averaged
3155.3 gms as compared to 3884.8 gms in GDM group.
Medication regimens were different between the two groups: while all 7PGD subjects required medication, only 2 GDM
subjects required medication. The level of diabetes control
between the PGD and GDM groups was different. For this
study, HbA1c≥7% (53 mmol/mol) was considered a marker
for poorly controlled disease. Accordingly, our data indicate
that 5 of the 7 PGD patients were poorly controlled, while all
of the GDM patients were well controlled. The minimum
diagnostic criteria based on HbA1c are 5.7% (39 mmol/mol)
for GDM and 6.5% (48 mmol/mol) for PGD. In our cohort, the
average HbA1c was 6.0% (42 mmol/mol) for GDM and 7.1%
(54 mmol/mol) for PGD subjects, showing an average increase
of only 0.053% above the diagnostic criteria for GDM, but an
increase of 0.09% above the diagnostic criteria for the PGD
group. Thus GDM group subjects had better controlled
disease than PGD subjects, possibly due to the shorter time
course of GDM as compared to PGD (Table 1).
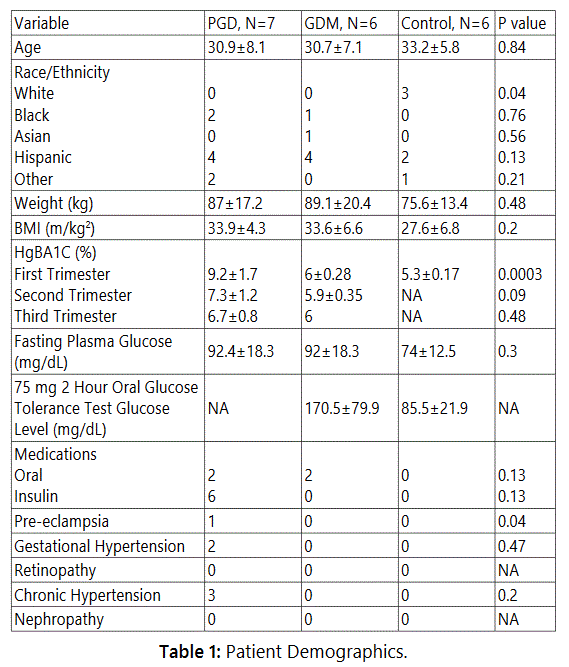
Average maternal BMI and neonate head circumference were similar between the GDM and PGD groups (Table 2). None of the PGD neonates in our cohort was macrosomic (average birth weight 3155.3±664.5 gms). Of the 6 GDM neonates, 3 were macrosomic (4485±86.07 gms) and 3 were normal (3285±337 gms). Thus phenotypically, both the mother and the newborn infant in the GDM group were different from that in the PGD group.
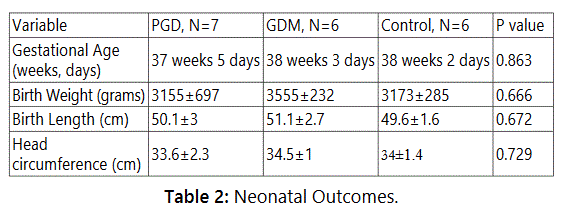
The Urinary Exosome Protein Content is Different in Diabetic versus Normal Pregnancy Subjects
Urine exosome proteomic data from an early time point of
pregnancy (24 hour urine samples from week 20) showed that
1103 proteins were identified with a spread of 645 CTRL
proteins, 855 GDM proteins and 733 PGD proteins in each
subject of the group.475 proteins were common to all three
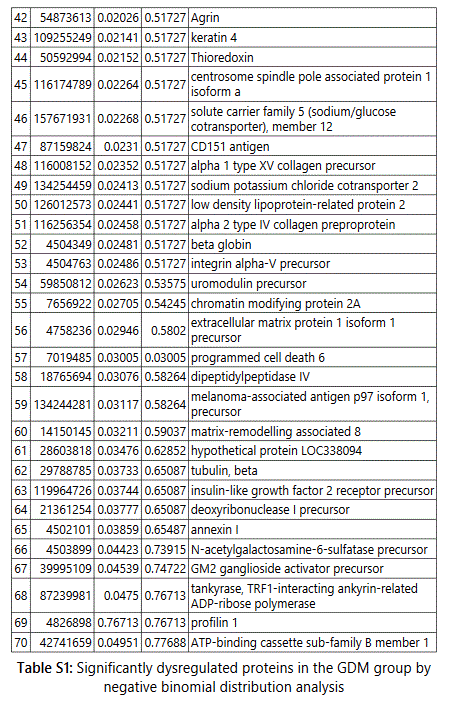
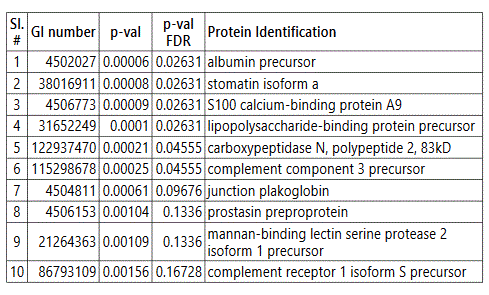
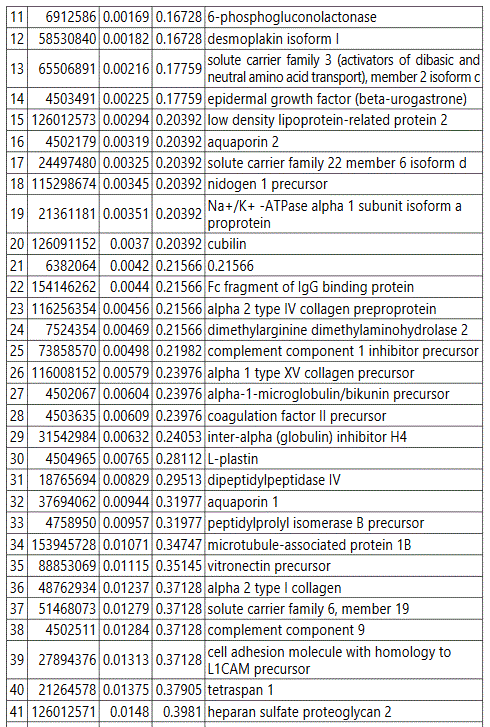
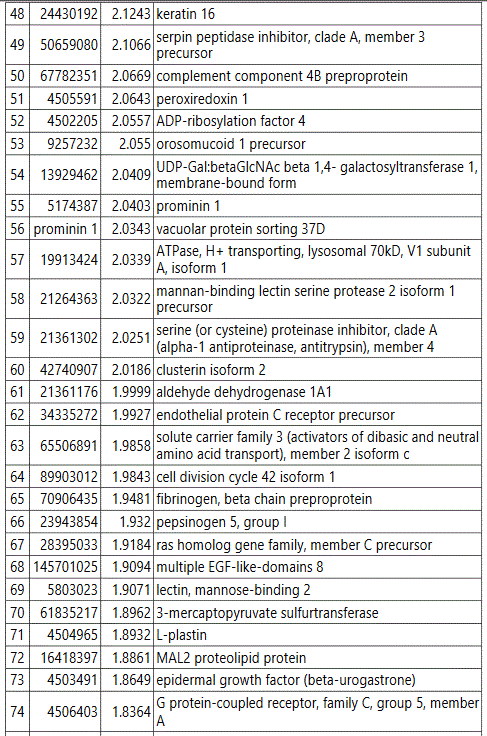
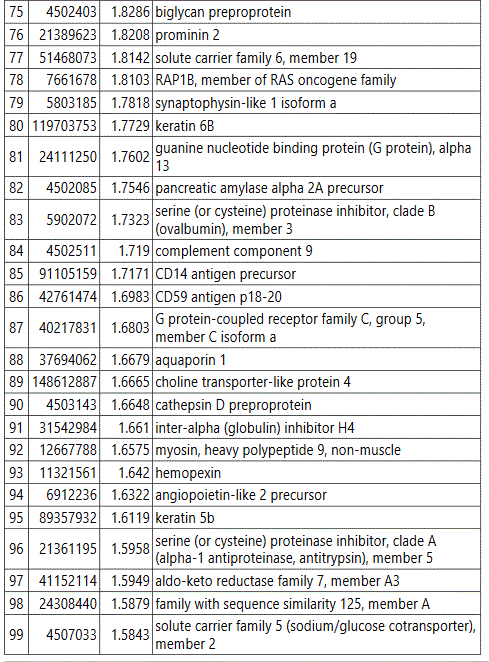
groups (Figure S1). Tables S1-S5 document the individual
protein identity of each of the groups. These protein data show
differences in the non-diabetic versus diabetic pregnancy urine
exosomes' protein signatures, and further differences between
GDM and PGD subjects at an early pregnancy time point.
To further delineate semi-quantitative differences between groups we used Negative Binomial Distribution to perform linear regression of the expression data of 1103 proteins. CTRL vs GDM analysis showed 70 proteins to be significantly different with S100 calcium-binding protein A9 (S100A9) the most different between the groups. (Table S1, FDR<10%). Similar analysis on CTRL vs PGD groups showed 77 proteins to be significantly different, with S100A9 as one of the 7 proteins reaching FDR<10% (Table S2).
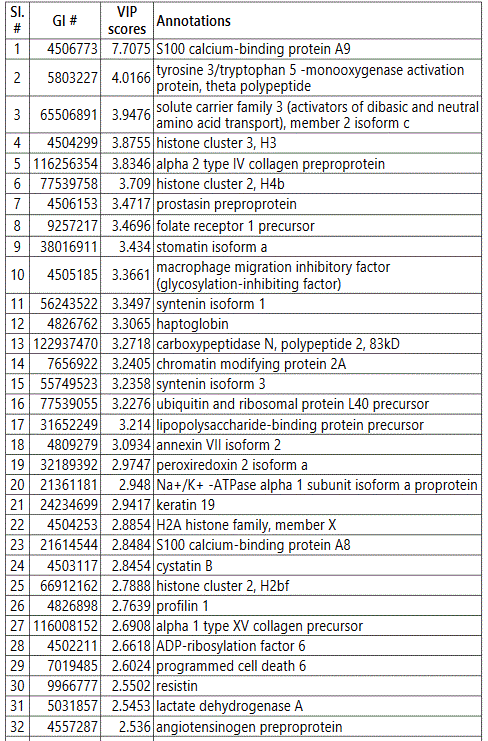
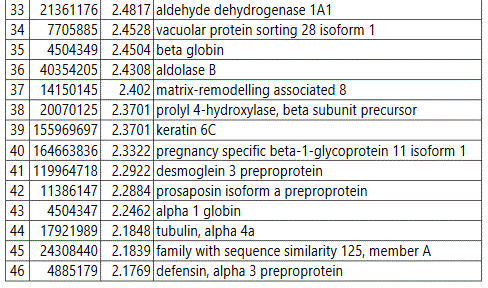
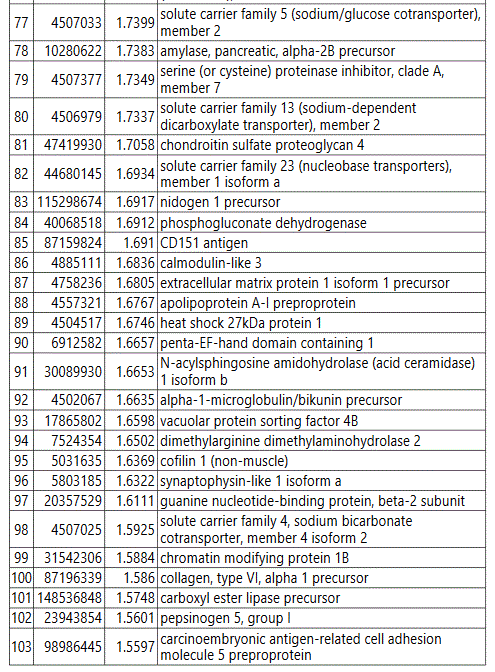
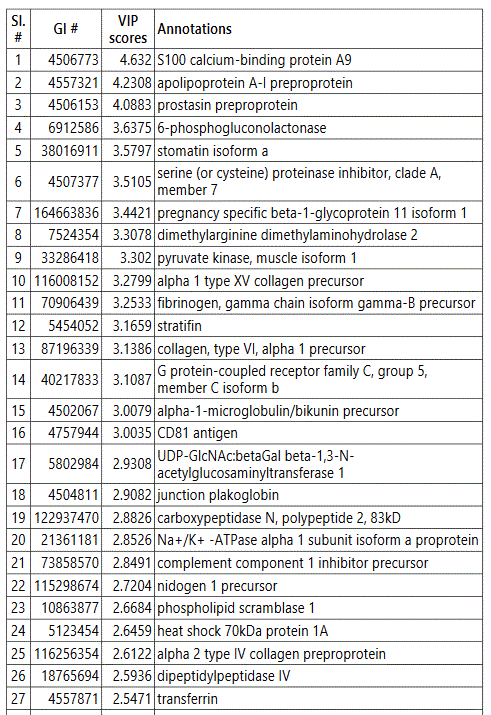
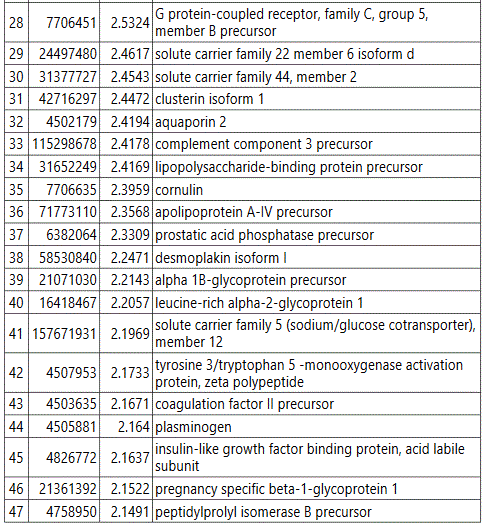
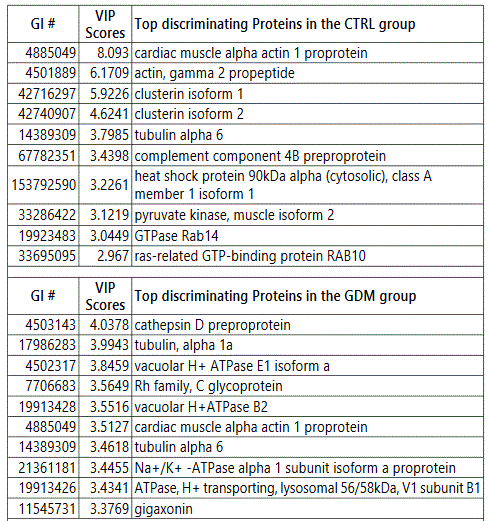
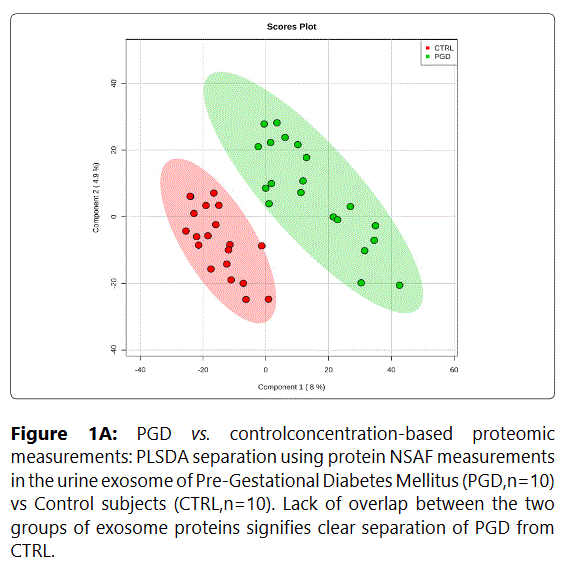
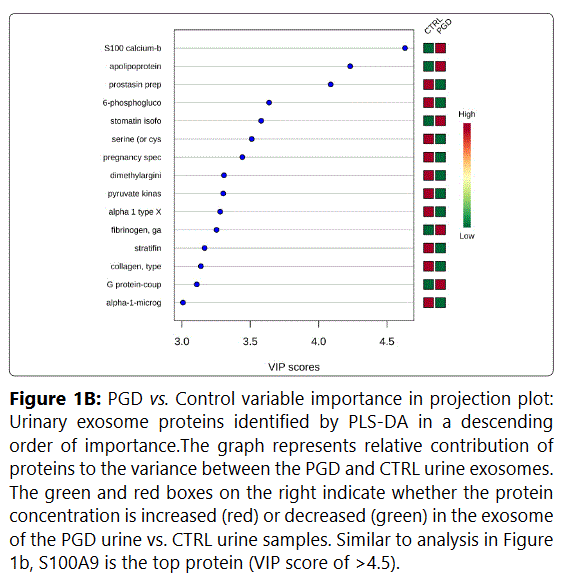
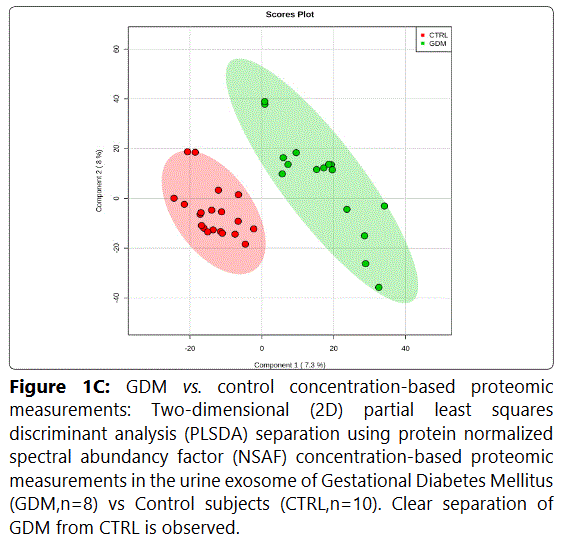
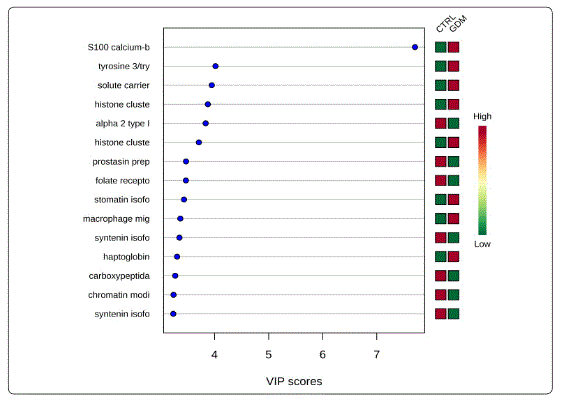
We used MetaboAnalyst suite to visualize the proteomic data. Partial Least Squares Discriminant (PLSD) analysis (Figures 1A-1C) showed a clear separation between CTRL and GDM proteins early in pregnancy. To determine which proteins contribute to this discrimination we conducted the Variable Importance in Projection (VIP) analysis (Figure 1D). Table S3 shows that 107 proteins contributed to this discrimination between CTRL and GDM groups, with a VIP score of >1.5, and S100A9 topped this list with a VIP score of 7.7.A similar analysis between CTRL and PGD groups showed clear distinction between protein expressions in the 2 groups and S100A9 with the top VIP score of 4.09 (Table S4).

S100A9 Protein Upregulation in Early Pregnancy Urine
Exosome of Diabetic Subjects is Validated by Western
Immunoblotting
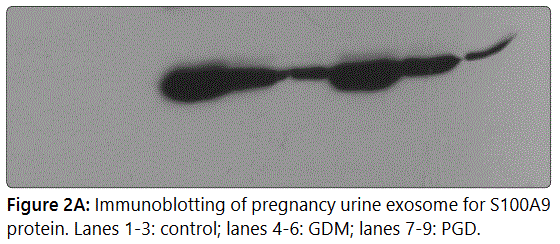
We validated the statistically significant upregulation of
S100A9 protein in diabetic urine exosomes by western
immunoblotting analysis. Equal amounts (50 μg) of protein
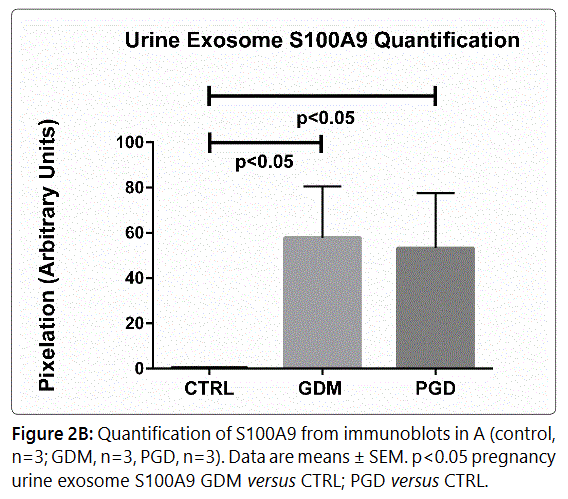
from each exosome sample was used, (Figure 2A, 2B) showing
significant S100A9 upregulation in diabetic exosome.
However, all diabetic patients did not uniformly show S100A9
upregulation. We therefore sought to further determine if the
demographic feature differences between subjects of the
same group correlate with the relative abundance of this
protein as denoted by its peptide count.
S100A9 Protein Peptide Count Data Correlates with Obesity
and Macrosomia Differently in GDM versus PGD subjects.
The average urinary exosome load of S100A9 peptide
count for the PGD group was 22.25±37.89 (N=10) while the
GDM had an average peptide count of 23.63±34.7 (N=8). Both were significantly different from the CTRL subject
peptide count of 1.15±2.39 (N=10) (p=0.0065, GDM vs CTRL
and p=0.0174, PGD vs CTRL), which also closely tracked with
the western blot data.
The average S100A9 peptide count was significantly different in the PGD and GDM subjects (7.4±13.43 versus 29.08±38.78, p=0.0209). When level of control of disease within the group was considered, PGD patients with well controlled disease have a lower average peptide count(0.25±0.5) compared with the poorly controlled group of PGD patients. It is noteworthy that both of these averages are less than the GDM group average.
GDM mothers with macrosomicneonates of had an average peptide count of 32.67±54.4 while GDM mothers of non-macrosomic neonates had an average peptide count of 25.5±17.85. Whereas this peptide count difference is likely less significant, it does illustrate a higher maternal peptide count for macrosomic neonates. Interestingly, a significantly higher rate of macrosomia in GDM than in PGD was observed (p=0.0002).
Lastly, we considered BMI. 6 of the 7 PGD patients and 4 of the 6 GDM patients were obese. Peptide count average of obese PGD patients was 6.25±12.44, while that of a single non-obese PGD patient was 0. In the GDM group, peptide average of obese patients was 42.5±37.53 and only 2.25±2.87 for non-obese patients. In the non-diabetic group, the peptide average of the obese subjects was 11.5±4.04 and only 1.67±1.51 for non-obese subjects. This strong trend (p=0.0585) demonstrates that peptide count is higher in pregnant obese women than in pregnant non-obese women.
Urine Exosome Protein Content Changes from Early to
Late Pregnancy Proceed via Different Pathways in CTRL
versus PGD versus GDM
Further, we sought to examine how progression of
pregnancy from an early time point to a later time point is
reflected in the urine exosome proteins and pathways in these
three groups. We performed a nested analysis on urine
exosomes from 4 CTRL, 5 GDM and 3 PGD subjects that
completed the 36th week collection in addition to the 20th
week samples.
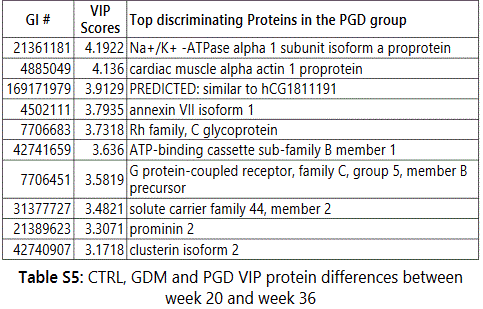
We observed considerable differences between exosome proteomes of early versus late pregnancy. For the 20th and 36th week samples respectively, 357 and 268 proteins (CTRL), 460 and 396 proteins (GDM) and 369 and 352 proteins (PGD) were identified. Thus the number of proteins identified in urine exosome generally decreased from early to late pregnancy. Groupwise analyses of the 20th and 36th week samples showed that the number of unique proteins identified reduced 5-fold in CTRL, roughly 2-fold in GDM and only slightly decreased in PGD groups (Figures S2A and S2B).
Thus, although the number of total pregnancy proteins identified was the highest in the GDM group (GDM>PGD>CTRL with 530,470 and 377 proteins respectively), within a group, the newer proteins identified during later pregnancy was CTRL < GDM < PGD with 20, 70 and 101 proteins respectively.
In addition, the protein pathways predominantly contributing to changes in progression of pregnancy from week 20 to week 36 were different in each group (Table S5, Figures S3A-S3C). We observed differences even in the extent to which a protein expression changed in one group versus the other (Δ1 =Protein1 NSAFWeek20 - Protein1 NSAFWeek36). Thus, Δ1 (for a given protein) for CTRL was different from Δ1 for GDM, which in turn was different from Δ1 for PGD group.
Discussion
Urine comprises at least two distinct exosome populations: exosomes that are produced by pre-glomerular organs such as brain and muscle that are filtered from plasma into urine, and those that are produced by the kidney tubular cells distal to glomerular filtration. Urine exosome proteins represent only 3% of the whole urine proteome [23] and their cellular nature allows for the strong likelihood that this 3% contains cell- and cell-state specific markers derived from both renal and pre-renal organs. Given that every living surface is shown to use these bio-active nanoparticles for intercellular communication, [24] urine exosomes potentially have a systemic level representation, and their analysis provides clinically relevant information in describing a disease phenotype such as diabetes during pregnancy. Two recent exosome proteomics analysis from (a) Pisitkun, et al. show 295 distinct proteins at least 22 of which have been implicated in various kidney and systemic abnormalities [25] and (b) Zhou, et al. show the utility of exosomes as a source of kidney physiology biomarkers in AKI subjects [26].
In this report, we examined the difference in urine exosome protein content of diabetic and non-diabetic pregnancy subjects. In all, we found that 1103 proteins were identifiable from a total of 28 urine exosome samples of non-diabetic and diabetic subjects. Using three independent methods, namely negative binomial distribution for linear regression analysis, partial least squares discriminant analysis, and variable importance in projection analysis of the proteomic data, we have shown that the calcium binding protein S100 A9 is significantly upregulated in both pre-gestational as well as gestational diabetic pregnancies. S100A9 upregulation in diabetics was validated by western immunoblotting. We also showed that the peptide number of this protein independently correlates with macrosomia in the newborn infant as well as maternal obesity, although to a different extent in the GDM versus PGD groups. Though the mechanism of S100A9 upregulation in diabetes/obesity and its clinical significance needs to be established, other recent studies [27-29] have demonstrated S100A9 upregulation in diabetes or obesity, further supporting our finding.
S100A9 upregulation as a biomarker of inflammatory processes and immune response is gaining fast ground. S100A9 heterotetramerizes with S100A8 to form calprotectin,which has a plethora of intracellular and extracellular functions.It is predominantly localized in the cytoplasm but is translocated to the cytoskeleton and cell membrane upon elevation of intracellular calcium level, and gets secreted to extracellular space to amplify innate immune response by acting as an endogenous Danger-Associated Molecular Pattern (DAMP) that promotes inflammation [30]. To our knowledge, this report is the first study showing S100A9 elevation in the urinary exosomes of human subjects with obese/diabetic pregnancy. Given its ability to induce neutrophil chemotaxis and adhesion, a role for S100A9 in exosome vesicle trafficking cannot be ruled out. If so, it would be interesting to address whether S100A9 would play a role in increased exosome production in diabetic milieu.
This study has numerous strengths. The 24-hour urine sample collection design lends itself to reduced variation and is more representative of the phenotype. Collection of samples at two time points from the same individuals allows for quantification of intra-subject variability. Additionally, by virtue of their lipid lamellar structure, exosomes serve to transport hydrophobic proteins, so the analysis is not limited to hydrophilic proteins only. One limitation of this study is its small sample sizes. The S100A9 upregulation should be confirmed in a larger population, with subjects that are more tightly grouped together and matched with respect to age, gravidity, and ethnicity. Additionally, the pre-gestational diabetic group did not include any patients with Type 1 diabetes, and therefore the results of this study are limited in their application to Type 1 diabetics and further studies in this population are needed. In our PGD population the date of diabetic diagnosis, and therefore duration of their disease, was not collected. As a result we were unable to analyze the data with regards to duration of disease. Further, the PGD vs GDM difference in S100A9 peptide count also needs to be verified in a larger population. It appears that when the data within the PGD group peptide count is considered, higher peptide count is associated with poorer diabetes control. Possibly by extending the time points of sample collection to include the 4th-8th week of pregnancy and 6 weeks after childbirth, this finding can be further validated. By developing a list of plausible risk-stratifiers, non-diabetic and/or non-obese pregnant women at risk for developing diabetes can potentially benefit from these studies.
Finally, we used 1d gel electrophoresis to resolve the exosome proteins prior to LC/MS-MS analysis, which resulted in identification of 1103 proteins overall. If we had conducted direct exosome protein trypsinization instead of following this method, perhaps the number of identified proteins might have increased. In our experience, exosomes cargo contains non-full length peptides as well, and has the potential to confound the analysis. By following the gel electrophoresis method, we ensured that we only compare full length protein differences between groups.
Conclusion
In this study, we summarize our initial findings from diabetic pregnancy urine exosome proteomic analyses. We used exosomes from 24 hour urine samples of diabetic pregnancy subjects obtained during the 20th week of pregnancy and compared them with exosomes from non-diabetic pregnancy subjects. Our data show major alteration to calcium-handling pathways and cellular-danger signal pathways between groups. The major finding of this study is that the DAMP protein S100A9 sorts into exosomes, and in diabetics this exosomal sorting is upregulated. The exosomal load of this protein correlates with not only maternal obesity but also macrosomia of the new born infant. Further, developmental changes from week 20 to week 36 in GDM and PGD phenotypes employ different protein pathways.
Given that maternal diabetes during human pregnancy alters the quality of life of both the mother and the newborn after childbirth, and considering the non-invasive nature of urinary exosome collection, processing and analysis of 24 hour urine samples is an attractive platform to study diabetes during pregnancy and its consequences.
Work from the Knepper group shows that many important renal proteins (e.g. aquaporins, polycystins and podocyn) are shed in the urine exosome. [25; 31] Our current report adds S100A9 to this group of functionally important proteins to be identified in urine exosomes. Our findings, together with other studies in pancreatic cancer tissue, saliva, and serum suggest that the upregulation of S100A9 DAMP signal is a common and valid biomarker of inflammatory processes and immune response. This report potentially forms the preliminary level basis for future studies wherein the pharmacologic and therapeutic potential of S100A9 and related pathways could be utilized to impart personalized care to a pregnant woman with diabetes.
Acknowledgements: The authors would like to thank Edith Esqueda for her efforts in helping to recruit patients to the study and provide the authors with clinic resources.
Conflicts of Interest: The author(s) report(s) no conflict of interest.
Funding: NIH NIDDK UAB/UCSD O'Brien Core Center for Acute Kidney Injury Research grant (NIH 1P30 DK 079337); the GSK-SSS funding for biomarkers; the UC San Diego CRCHD (NIH/NIMHD-P60 MD000220); and the Proteomics Catalyst grant instituted by UC San Diego Clinical Translational Research Institute.
References
Supplementary Data