5th International Conference on Oncology & Virology
July 25-26, 2019 | Holiday Inn Rome Aurelia, Rome, Italy
Risk Stratification of Cervical Lesions Using Capture Sequencing and Machine Learning Method Based on HPV and Human Integrated Genomic Profiles
1Department of Obstetrics and Gynecology, The First Affiliated Hospital of Sun Yat-Sen University, China
2Department of Neurology, The First Affiliated Hospital of Sun Yat-Sen University, China
Background: From initial HPV infection and precursor stages, the development of cervical cancer takes decades. High-sensitivity HPV DNA testing is currently recommended as primary screening method for cervical cancer, while better triage methodologies are encouraged to provide accurate risk management for HPV positive women.
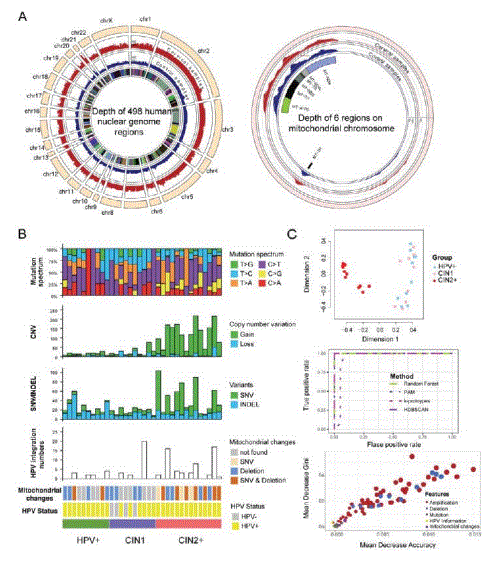
Methods: Given that virus-driven genomic variation accumulates during cervical carcinogenesis, we designed a 39 Mb custom capture panel targeting 17 HPV types and 522 mutant genes related to cervical cancer. Using capture-based next-generation sequencing, HPV integration status, somatic mutation and copy number variation were analyzed on 34 paired samples, including 10 cases of HPV infection (HPV+), 10 cases of CIN1 and 14 cases of CIN2+ (CIN2: n=1; CIN2-3: n=3; CIN3: n=9; SCC: n=1). Finally, the Machine Learning Algorithm-Random Forest was applied to build the risk stratification model for cervical precursor lesions based on CIN2+ enriched biomarkers.
Results: Generally, HPV integration events (11 in HPV+, 25 in CIN1 and 56 in CIN2+), non-synonymous mutations (2 in CIN1, 12 in CIN2+) and copy number variations (19.1 in HPV+, 29.4 in CIN1 and 127 in CIN2+) increased from HPV+ to CIN2+. Interestingly, “common” deletion of mitochondrial chromosome was significantly observed in CIN2+ (P value=0.009). Together, CIN2+ enriched biomarkers, classified as HPV information, Mutation, Amplification, Deletion and mitochondrial change, successfully predicted CIN2+ with average accuracy probability score of 0.814, and Amplification and Deletion ranked as the most important features.
Conclusion: Our custom capture sequencing combined with machine learning method effectively stratified the risk of cervical lesions and provided valuable integrated triage strategies.
Biography
Zifeng Cui is a PhD student. Presently working under Professor Zheng Hu (Doctoral Supervisor, Chief Special Scientist of the Ministry of Science and Technology, Young Top Talents in Ten-thousand Talents Program)


